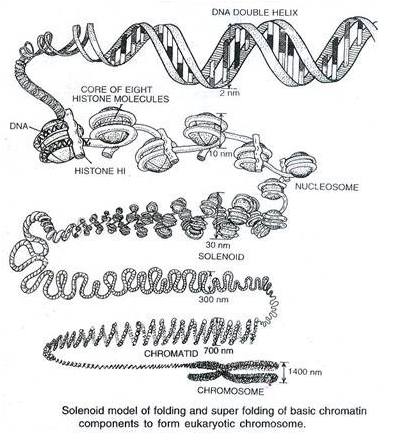
Solenoid model
The long sized DNA are accommodated in small areas (about 1 μm in E. coli and 5 μm nucleus in human beings) only through packing or compaction. The packaging of DNA in nucleosome is explained with the solenoid model. DNA is acidic due to the presence of a large number of phosphate groups. Compaction occurs by folding and attachment of DNA with basic proteins, non-histone in prokaryotes and histones in eukaryotes.

The packaging of DNA in nucleosome is carried out with help of histones. Out of five types of histone proteins, four (H2A, H2B, H3 and H4) occur in pairs to produce histone octamer, called nu body or core of nucleosome. Their positively charged ends (due to basic amino acids) are towards the outside. They attract negatively charged strands of DNA. About 166 bp of DNA is wrapped over nu body for 1% turns to form nucleosome of size 110 x 60A (11×6 nm). DNA connecting two adjacent nucleosomes is called interbead or linker DNA. It bears H 1 histone protein (called plugging protein and act as marker protein). Length of linker DNA is varied (about 145A with 70 bp). Nucleosome and linker DNA together constitute chromatosome.
Nucleosome chain gives a beads on string appearance under the electron microscope. The beaded string is coiled to form cylindrical coil or solenoid having 6 nucleosomes per turn. Actually the nucleosomal organisation has approximately 10nm thickness, which gets further condensed and coiled to produce a solenoid of a 30nm diameter. This solenoid structure undergoes further coiling to produce a chromatin fibre of 30-80 nm and then a chromatid of 700 nm.
DNA (2 nm diameter) → nucleosome (10 nm diameter) → solenoid (30 nm diameter) → chromatin fibre (30-80 nm diameter) → chromatid (700 nm diameter) → chromosome (1400 nm diameter)
Chromatin is held over a scaffold of nonhistone chromosomal or NHC proteins. At some places chromatin is densely packed to form darkly staining heterochromatin. At other places chromatin is loosely packed. It is called euchromatin. Euchromatin is lightly stained.