MUTANT DETECTION
When collecting mutant of a particular organism, one must know the normal or wild type characteristics so as to recognize an altered phenotype. Usually, mutation are rare, about one per 107 to 1011 cells, therefore, it is necessary to have a very sensitive direction system and one has to look at perhaps thousands to millions of colonies or clones. Mutation detection systems in bacteria and other haploid organism are straight forward, because any new allele should be seen immediately, if it is a recessive mutation.
Sometimes detection of mutants is direct of albino mutants of a normally pigmented bacterium are being studied, detection simply requires visual observation of colony colour.
Other direct detection systems are more complex, for example, the replicate plating technique is used to detect auxotrophic mutants. It distinguish between mutants and the wild type strain based on their ability to grow in the absence for a particular biosynthetic end product.
MUTANT SELECTION
1. Selection Technique: An effective selection technique uses incubation conditions under which the mutant will grow, because of properties given to it by the mutation, whereas the wild type will not. It involves reversions, mutations or the development of resistance to environmental stress.
2. Resistance Selection Technique: It follows a similar approach often wild type cells are not resistant to phage attack or antibiotic treatment, so it is possible to grow the bacterium in the presence of the agent and took for surviving organisms. Consider the example of a phage sensitive wild type bacterium when the organism is cultured in medium lacking the virus and then plated out on selective medium containing phages, any colonies that form will be resistant to phage attack and very likely will be mutants in this regard. Resistance selection can be used together with virtually any environmental parameter. Resistance to bacteriophage antibiotics or temperature are most commonly employed.
3. Substrate Utilization Technique: This technique is also employed in bacterial selection. Many bacteria catabolize only a few primary carbon source, with these bacteria, it is possible to select mutants by a method similar to that used in resistance selection. The culture is plated on medium containing an alternate carbon source. Any colonies that appear can use the substrate and are probably mutants.
4. Mutant Detection and Selection Methods: These methods are used for purposes other than understanding more about the nature of genes or the biochemistry of a particular micro-organism. Replica plating used for the isolation of auxotroph mutants.
CLASSIFICATION OF AUXOTROPHS
The classification of mutants of organisms can be done by replica plating technique. Here a suspension of bacterial cells treated with mutants is poured onto a master plate containing medium supplemented with those growth factors which will allow the desired mutants to grow. After incubation, the replication is applied to the plate and this picks up few colonies from the master plate, and it is pressed successively on series of plates containing minimal medium along and with various supplements. After further incubation, colonies appear on these replica plates in the same pattern as on the master plate, absence of a particular colony from a plate is an evidence that it requires that supplement missing from the plate, and so that mutant is classified according to the missing growth factor, e.g., arginine requiring mutants, threonine requiring mutants, etc.
Auxotrophic Mutants
Many variants of bacteria have been isolated which are unable to synthesize one or more amino acids or other factors required for growth. Such strains are called auxotrophs, while nutritionally independent, wild-type strains are called prototrophs. In E. coli strains with lac’ are prototrophs and those with lac are auxotrophs. An auxotrophic strain may be identified by its inability to grow and multiply in a synthetic medium lacking its specific growth requirements, in which the wild-type, prototrophic strain flourishes. If the synthetic medium is supplemented with the growth factor which the auxotroph cannot synthesize for itself, then auxotrophs and protrophs can grow equally well in it and can be indistinguishable. Similarly, strains of bacteria which have become resistant to the lethal action of antibiotics or bacterial viruses may appear identical to the wild type unless the drug or the phage is present in the environment to detect the difference between them. Usually, auxotrophs are given the symbol (-) and prototrophs the symbol (+).
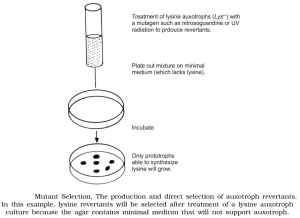
Auxotroph Selection
It is easy to pick out a prototrophic mutant from a mixture of auxotrophs and prototrophs because practically auxotrophs will not grow in a selective medium not containing the missing growth factor. But it is difficult to pick out an auxotroph from a medium because the prototrophs will grow pretty as well as the auxotrophs when the medium is supplemented.
We can solve this problem by the fact that penicillin kills only growing bacteria. If a suspension of E. coli prototrophs is treated with a mutagen and then grown in a minimal medium containing a lethal concentration of penicillin, all the cells which are capable of growth will be killed. After a suitable delay, the bacteria remaining including any auxotrophs present which cannot grow because of lack of supplements are centrifuged down, washed, re-suspended in a penicillin-free medium, and poured on supplemented agar plates. The resultant colonies are tested and classified as requiring various amino acids or vitamins for normal growth.