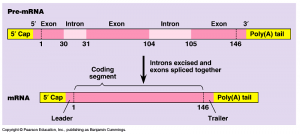
Post-transcriptional modification is a process in cell biology by which, in eukaryotic cells, primary transcript RNA is converted into mature RNA. A notable example is the conversion of precursor messenger RNA into mature messenger RNA (mRNA), which includes splicing and occurs prior to protein synthesis. This process is vital for the correct translation of the genomes of eukaryotes as the human primary RNA transcript that is produced as a result of transcription contains both exons, which are coding sections of the primary RNA transcript and introns, which are the non coding sections of the primary RNA transcript.
mRNA processing
The pre-mRNA molecule undergoes three main modifications. These modifications are 5′ capping, 3′ polyadenylation, and RNA splicing, which occur in the cell nucleus before the RNA is translated.
a) 5′ Processing
Capping. Capping of the pre-mRNA involves the addition of 7-methylguanosine (m7G) to the 5′ end. To achieve this, the terminal 5′ phosphate requires removal, which is done with the aid of a phosphatase enzyme. The enzyme guanosyl transferase then catalyses the reaction, which produces the diphosphate 5′ end. The diphosphate 5′ prime end then attacks the α phosphorus atom of a GTP molecule in order to add the guanine residue in a 5’5′ triphosphate link. The enzyme (guanine-N7-)-methyltransferase (“cap MTase”) transfers a methyl group from S-adenosyl methionine to the guanine ring. This type of cap, with just the (m7G) in position is called a cap 0 structure. The ribose of the adjacent nucleotide may also be methylated to give a cap 1. Methylation of nucleotides downstream of the RNA molecule produce cap 2, cap 3 structures and so on. In these cases the methyl groups are added to the 2′ OH groups of the ribose sugar. The cap protects the 5′ end of the primary RNA transcript from attack by ribonucleases that have specificity to the 3’5′ phosphodiester bonds.

b) 3′ Processing
Cleavage and Polyadenylation. The pre-mRNA processing at the 3′ end of the RNA molecule involves cleavage of its 3′ end and then the addition of about 200 adenine residues to form a poly (A) tail. The cleavage and adenylation reactions occur if a polyadenylation signal sequence (5′- AAUAAA-3′) is located near the 3′ end of the pre-mRNA molecule, which is followed by another sequence, which is usually (5′-CA-3′). The second signal is the site of cleavage. A GU-rich sequence is also usually present further downstream on the pre-mRNA molecule. After the synthesis of the sequence elements, two multisubunit proteins called cleavage and polyadenylation specificity factor (CPSF) and cleavage stimulation factor (CStF) are transferred from RNA Polymerase II to the RNA molecule. The two factors bind to the sequence elements. A protein complex forms that contains additional cleavage factors and the enzyme Polyadenylate Polymerase (PAP). This complex cleaves the RNA between the polyadenylation sequence and the GU-rich sequence at the cleavage site marked by the (5′-CA-3′) sequences. Poly (A) polymerase then adds about 200 adenine units to the new 3′ end of the RNA molecule using ATP as a precursor. As the poly (A) tails is synthesised, it binds multiple copies of poly(A) binding protein, which protects the 3’end from ribonuclease digestion.
c) Splicing
RNA splicing is the process by which introns, regions of RNA that do not code for protein, are removed from the pre-mRNA and the remaining exons connected to re-form a single continuous molecule. Although most RNA splicing occurs after the complete synthesis and end-capping of the pre-mRNA, transcripts with many exons can be spliced co-transcriptionally. The splicing reaction is catalyzed by a large protein complex called the spliceosome assembled from proteins and small nuclear RNA molecules that recognize splice sites in the pre-mRNA sequence. Many pre-mRNAs, including those encoding antibodies, can be spliced in multiple ways to produce different mature mRNAs that encode different protein sequences. This process is known as alternative splicing, and allows production of a large variety of proteins from a limited amount of DNA.