Plasmid
Plasmids are small circular double-stranded, chromosomal DNA molecules, which occur in most bacterial organisms. The term was first introduced by Joshua Lederberg in 1952. Plasmids are separated and replicated separately from the bacterial genome. They usually only bear a few chromosomes. In bacteria and certain archaea, plasmids are present. The plasmid is not an important bacterial element but is often advantageous to the bacteria. Plasmids have a high copy number. The size varies from 1 to 1000kbp. Plasmids serve as important tools in genetics and biotechnology labs, where they are commonly used to multiply (make many copies of) or express particular genes. Many plasmids are commercially available for such uses. The gene to be replicated is inserted into copies of a plasmid containing genes that make cells resistant to particular antibiotics and a multiple cloning site (MCS, or polylinker), which is a short region containing several commonly used restriction sites allowing the easy insertion of DNA fragments at this location.
Features of Ideal Cloning Vector
- Small size
- Origin of replication
- Low molecular weight
- Must be easier to handle
- Resistant to stress
- Easier to isolate and transform into bacteria
- Ability to confer selectable traits
- Easier to screen
- Must have multiple cloning sites with a large number of restriction sites
Let’s discuss why plasmid is an important cloning vector!
In genetic engineering, we need gene of interest copies in numerous numbers. So we use a vector, mostly a plasmid to multiply the gene of interest. It is done by incorporating the vector inside a bacteria and make multiple copies. As the bacteria grow in optimal temperature and Ph conditions, the plasmids that hold the gene of interest also multiply. Plasmid vectors are important to have multiple cloning sites with various restriction enzyme sites, they have selectable marker genes, and they have a way to detect when a foreign piece of DNA.
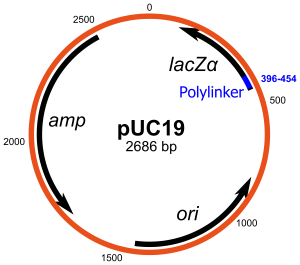
Origin of Replication
The ori is a sequence in which DNA replication starts enabling a plasmid to reproduce itself such that it has to thrive within cells. Some replication origins produce several copies of plasmid, and others only produce few copies, based on how they are regulated.
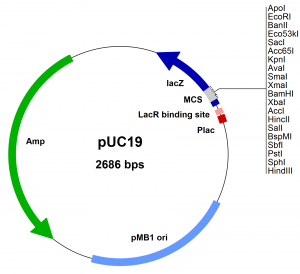
Multiple cloning sites
A multiple cloning site (MCS) is a DNA region within a plasmid containing multiple specific sites for the enzyme restriction cutting. Without affecting all other plasmids it permits the introduction of foreign DNA.
Steps Involved in Cloning

Selectable marker
A selectable marker may be used to classify and erase non-transformers and to selectively expand processors. A selectable marker is a gene injected into a cell that confers a suitable artificial selection characteristic, in particular a bacterium or cultivated cell. Selectable marker genes can be divided into genes that impart toxic compounds such as antibiotics that destroy or otherwise compromise untransformed tissue that can be produced by their development. During transformation, the bacteria have chances of degrading the plasmid or not taking up the plasmid. So adding up the selectable marker would distinguish the bacteria having the plasmid. Mostly there is the incorporation of antibiotic resistance gene that is encoded by the plasmid. So that the bacteria which have the antibiotic resistance gene would only grow in the media.
Plasmids are to make large amounts of proteins. In this case, researchers grow bacteria containing a plasmid harboring the gene of interest. Just as the bacterium produces proteins to confer its antibiotic resistance, it can also be induced to produce large amounts of proteins from the inserted gene. This is a cheap and easy way of mass-producing a gene or the protein it then codes for, for example, insulin or even antibiotics.
Screening Markers
There are chances of bacteria having self-ligation and not the gene of interest. So screening markers are used majorly for dying it. The colored culture would distinguish the bacteria having a gene of interest.
Extraction of Gene of Interest
The extraction of the gene of interest is done by using the same restriction enzyme present in the plasmid. The same restriction enzyme cleaves the plasmid. The DNA ligase then hybridizes the gene of interest in the plasmid to produce a recombinant DNA molecule. Then the transformation process continues to transfer the plasmid into the bacterial cell. The bacteria having a gene of interest or the one which shows the colored culture can then be produced in bulk amounts.
Integration of Plasmid into the chromosome
Instead of the integrative plasmid with the homologous recombination design, most of the bacteria cannot be converted with linear DNA. Microorganisms with replicative plasmids may be supplemented with DNA, but these are intrinsically unstable. In order for exogenous DNA to be stabilized, a gene must be irreversibly incorporated within the cell into a functional molecule of DNA. This can be achieved by a homologous process of recombination.
- Recombination- Plasmid recombine with chromosomes when they both share a common sequence ie. Homologous sequence.
- Transposition- Plasmids insert into chromosomes by transposons and forms a high frequency of recombination (The plasmid entry into the chromosome is through the homologous recombination.) Transposition may further facilitate other forms, including deletions, reversions, and replicon fusions, of DNA rearrangements. Such DNA rearrangements may have a deep impact. It will result in new genetic material being obtained stably.
Limitations using the Plasmid Vector
- The maximum limit for cloning DNA is 12kb.
- Preparation of competent host cells is necessary.
- Not efficient for generating genomic libraries as overlapping regions needed to be in the proper sequence.
- Eukaryotic protein expression is mostly not possible in this vector as it only solves the purpose for cloning vector and not expression vector.
References
https://www.bocascientific.com/index.php?main_page=popup_image&pID=774https://commons.wikimedia.org/wiki/File:PUC19.svghttps://www.sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology/selectable-markerhttps://blog.addgene.org/plasmid-101-origin-of-replicationhttps://www.ncbi.nlm.nih.gov/books/NBK21498/figure/A1585/?report=objectonlyhttps://www.ncbi.nlm.nih.gov/pmc/articles/PMC3333862/#:~:text=As%20plasmids%20are%20circular%2C%20a,to%20revert%20to%20wild%2Dtype.https://www.ncbi.nlm.nih.gov/books/NBK21498/https://www.sciencedirect.com/topics/neuroscience/cloning-vector
