Isolation of Genetic Materials
The knowledge of gene isolation was developed after gaining the concept of physical and chemical characteristics of described DNA fragments their sizes, shapes and conformation can aid in the selection of methods used to isolate and purify those segments.
Plants provide several special challenges for researchers interested in the recombinant DNA research. No other class of living organisms has three separate genomes to analyze a nuclear genome containing high molecular weight linear chromosomes, a circular mitochondrial genome, and a circular chloroplast genome. Intact, high molecular weight (>150kbp) plant nuclear gDNA is used to construct genomic DNA libraries and to probe the plant genome for the presence of DNA markers, such randomly amplified polymorphic DNAs (RAPDs) and restriction fragment length polymorphisms (RFLPs). Chloroplast cpDNA and mitochondrial mtDNA, which are thought to incur fewer structural changes over time, are often used to study plant systematic. They are also used to study in vitro synthesis and assembly of organelle proteins.
Isolating the DNA (Gene) for interest
For isolation of DNA from linear chromosomes of living organisms like plants, animals or from simple bacteria, it is necessary to break the cell for release of DNA which may be done by physical (homogenization or pressure), or biochemical means like solubilization with detergents or enzymes. The selection of enzyme for this process is dependent on the composition of the Particular cells that are used for plant materials, for fungal sample, chitinase is used.
Lysozyme is commonly used for both bacterial and animal tissue samples. The DNA so obtained is then separated from the cell walls, membranes, and other cellular debris by means of centrifugation, often followed by density gradient centrifugation. The localization of DNAs is the gradient may be done by instrumental method (absorption at 260 nm or visually by means of fluorescence). The isolation of DNA is done by inserting a syringe into the side of the soft plastic centrifuge tube and withdrawing the bands visible in UV radiation. The isolated DNA may be further purified and physically characterized (for example, for size) by means of electrophoresis in gel made up of agarose or of acryl amide.
Isolation of genes is the in vitro gene manipulation as it gives us the ability to bypass the multiplicity of mechanisms that restrict gene transfer between unrelated organisms and are far more accurate and subtle in nature. The broad outlines of this technique is: isolation of gene of interest, insertion of a gene into an appropriate vector, transfer of foreign DNA (gene) through the vector into an appropriate host organism to produce recombinant DNA and identification of recombinants.
DNA Isolation Methods
Many different methods and technologies are available for the isolation of genomic DNA. In general, all methods involve disruption and lysis of the starting material followed by the removal of proteins and other contaminants and finally recovery of the DNA. Removal of proteins is typically achieved by digestion with proteinase K, followed by salting-out, organic extraction, or binding of the DNA to solid-phase support (either anion-exchange or silica technology).DNA is usually recovered by precipitation using ethanol or isopropanol. The choice of a method depends on many factors: the required quantity and molecular weight of the DNA, the purity required for downstream applications, and the time and expense.
The separation of DNA from cellular components can be divided into four stages:
- Disruption
- Lysis
- Removal of proteins and contaminants
- Recovery of DNA
Preparation of crude lysates
An easy technique for isolation of genomic DNA is to incubate cell lysates at high temperatures (e.g., 90°C for 20 minutes), or to perform a proteinase K digestion, and then use the lysates directly in downstream applications. Considered “quick-and-dirty” techniques, these methods are only appropriate for a limited range of applications. The treated lysate usually contains enzyme inhibiting contaminants, such as salts, and DNA is often not at optimal pH. Furthermore, incomplete inactivation of proteinase K can result in false negative results and high failure rates. It is not recommended to store DNA prepared using this method, as the high levels of contamination often result in DNA degradation.
Salting-out methods
Starting with a crude lysate, ”salting-out” is another conventional technique where proteins and other contaminants are precipitated from the cell lysate using high concentrations of salt such as potassium acetate or ammonium acetate. The precipitates are removed by centrifugation, and the DNA is recovered by alcohol precipitation. Removal of proteins and other contaminants using this method may be inefficient, and RNase treatment, dialysis, and/or repeated alcohol precipitation are often necessary before the DNA can be used in downstream applications. DNA yield and purity are highly variable using this method.

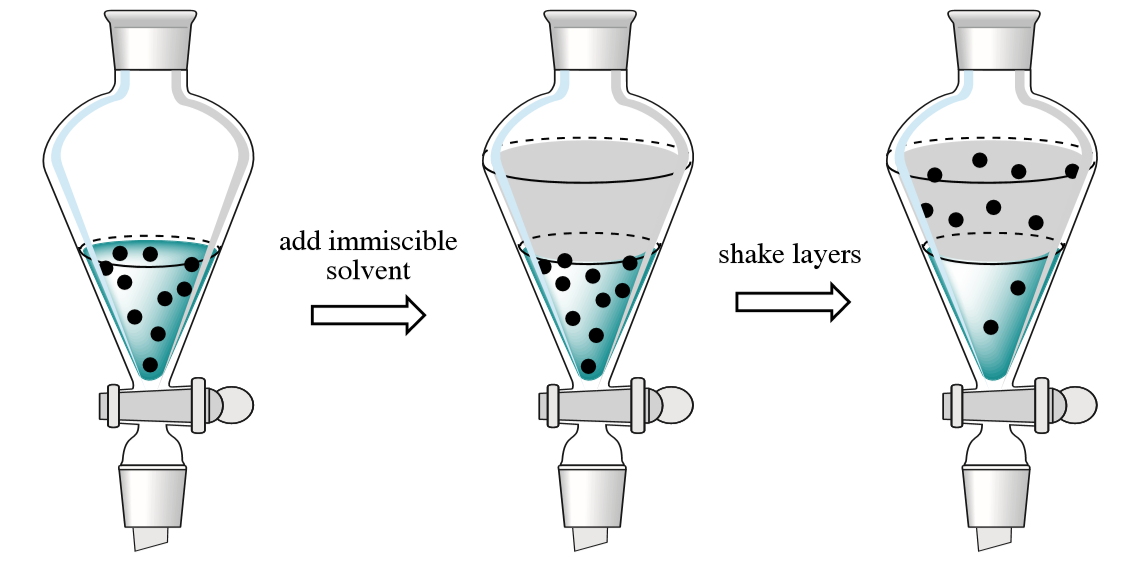
Organic extraction methods
Organic extraction is a conventional technique that uses organic solvents to extract contaminants from cell lysates. The cells are lysed using a detergent, and then mixed with phenol, chloroform, and isoamyl alcohol. The correct salt concentration and pH must be used during extraction to ensure that contaminants are separated into the organic phase and that DNA remains in the aqueous phase. DNA is usually recovered from the aqueous phase by alcohol precipitation. This is a time consuming and cumbersome technique. Furthermore, the procedure uses toxic compounds and may not give reproducible yields. DNA isolated using this method may contain residual phenol and/or chloroform, which can inhibit enzyme reactions in downstream applications, and therefore may not be sufficiently pure for sensitive downstream applications such as PCR. The process also generates toxic waste that must be disposed of with care and in accordance with hazardous waste guidelines. In addition, this technique is almost impossible to automate, making it unsuitable for high-throughput applications.
Cesium chloride density gradients
Genomic DNA can be purified by centrifugation through a cesium chloride (CsCl) density gradient. Cells are lysed using a detergent, and the lysate is alcohol precipitated. Resuspended DNA is mixed with CsCl and ethidium bromide and centrifuged for several hours. The DNA band is collected from the centrifuge tube, extracted with isopropanol to remove the ethidium bromide, and then precipitated with ethanol to recover the DNA. This method allows the isolation of high-quality DNA, but is time consuming, labor intensive, and expensive (an ultracentrifuge is required), making it inappropriate for routine use. This method uses toxic chemicals and is also impossible to automate.
Fragmentation by Mechanical Shearing:
Random fragments of DNA can be generated by mechanical shearing. The desired fragments obtained by this method are without cohesive ends; therefore, the ligation (joining) with the vector can be facilitated with a process known as homopolymer tailing. For example, a tail of dC residues (deoxynucleotide triphosphate) can be added to 3-OH terminus of DNA of sheared fragment and a tail of dG residues vector. The fragments to be cloned can then be joined to the reactor by annealing the tails.
Short- Gun method:
There is a restriction enzyme which is specific for a six-base sequence of DNA. It is used to cut foreign DNA to get a piece of interest. One can obtained fragments of 3 to 4 genes by this method, if distribution of bases is random. The fragments obtained can be too small or too big if the no of cutting sites recognized by enzyme used are too frequent and too sparse in the distribution. It is also important that is cutting sites falls on either end of gene of interest and not in between. When the same enzyme is used to cut the vector DNA, cohesive ends are created, in most cases depending on the nature of enzyme, the joining of vector and the isolated DNA is possible.

cDNA method:
If animal gene is to be expressed in a bacterium or yeast, a suitable method is to isolate the messenger RNA (mRNA), first, concerned with the specific protein production. It is often impossible to express animal gene directly in bacteria because of the introns within the gene. The protein encoding parts of the genes are exons. Mature mRNA molecules in animals cells do not contain sequences complementary to introns as they are removed by processing. Introns are the sequences in DNAs which are in fact transcribed into mRNAs but are subsequently split out are therefore form no permanent consequents of the mRNAs (only reverse transcriptase and DNA polymerase. Thus cDNA produced for a particular product can be then joined to the appropriate vector for cloning by homopolymer tailing as described above or proper linkers.
