Regulation of miRNA
miRNA: Introduction and History
- RNA (RNAi) is a biological mechanism in which gene expression is inhibited by the inhibition of RNA molecules usually by disrupting certain mRNA molecules.
- Other names, including cosuppression, post-transcriptional gene silencing (PTGS), and quelling, were historically known. For their work on the RNA interference in the nematode worm Caenorhabditiseligans, which they released in 1998, Andrew Fire and Craig C. Mello shared the 2006 Nobel Prize in Physiology or Medicine.
- miRNAs (microRNAs) are short non-coding RNAs that posttranscriptionally regulate gene expression.
miRNA Function
- MIRNA typically binds to the 3′-UTR of the target mRNAs and suppresses the development of proteins by destabilizing the mRNA and translational silencing.
- The RISC complex is oriented towards the mRNA to be degraded after biogenesis of the miRNA and the development of RISC.
- The miRNA loaded into the complex aims at RISC unique binding sites in the 3′′ untranslated mRNA transcript (UTR) region through base-pairing interactions in the seed sequence of the maturity of the miRNA:mRNA sequence, which is 7- to the 8-nucleotide sequence.
- For miRNA:mRNA and miRNA-mediated gene silence, the RISC is a ribonucleoprotein complex.
- Different proteins, including argonaute protein (AGO1 to AGO4) and Dicer, have been identified as RISC components.
- The Argonautic protein is a significant component of endonucleolytic cleavage in the RISC.
- Dicer functions as a ribonuclease in miRNA biogenesis leading to the release of the mature miRNA duplex.
Regulation of miRNA
The regulatory RNAs regulate gene expression by following two mechanisms:
- Degradation of mRNA
- Translation Arrest
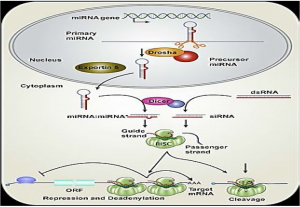
Degradation of mRNA
- It is well known how to control translations by degrading the mRNA sequence by small RNA. The RISC with a bonded siRNA recognizes and degrades additional RNA messenger (mRNA) molecules, resulting in dramatically reduced protein translation and efficient gene shutting.
- As previously mentioned, siRNA are short RNA sequences (~21 nt) made from long double-stranded RNA precursors, unlike miRNA, which is produced from the precursors encoded by the genome.
- The siRNA needs to be degraded with sequences that are identical to the mRNA. This widespread homology across the entire siRNA:mRNA sequence results in post-transcription control of gene expression by the use of components from miRNA machinery, like Dicer, to cleavage the target mRNA sequence at the binding site.
- Endonucleases, called Argonaute Proteins, are the active components of the induced RNA silencing complex (RISC), which complement their binding siRNA, by cleaving the target mRNA chain.
- As the fragments produced by dicer are double-stranded, they could each in theory produce a functional siRNA.
- However, only one of the two strands known as the lead strand binds the protein of argonaute and directs the silencing of genes.
- During RISC activation, the other anti-guide strand is degraded. The hexophane of strands is ATP-independent and conducted directly by components of the proteins of RISC.
Translation Arrest
The miRNA controls translation by multiple mechanisms:
(i) Deadenylation:
Translation Repression by miRNA is triggered by deadenylation of the poly-A tail and then mRNA sequence decapping. As a result, the mRNA sequence becomes unstable and is degradable. The abundance of mRNA is diminished and the translation reduces thereafter. Proof implies miRNA.
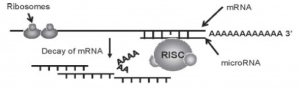
(ii) Blocking of initiation:
The miRNA binds to the RISC and attacked the 3’UTR of the mRNA sequence similar to the previously mentioned method. However the miRNA: RISC complex to the 3’UTR is intended to activate an event series that prevents initiation proteins from binding to the 5′ cap of mRNA rather than to induce deadenylation to trigger declines in mRNA.
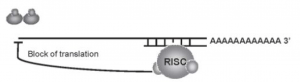

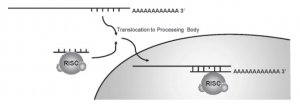
(iii) Translocation to Processing Bodies:
A third possible mechanism of miRNA action involves posttranscriptional regulation after the miRNA: RISC complex binds the mRNA target by translocating the miRNA:mRNA complex to cytoplasmic foci in the cell, referred to as processing bodies (P-bodies). P bodies do not contain translation ribosomal proteins but have enzymes and translation repression and mRNA turnover factors.
Applications of RNAi
- Gene Knockdown: In experimental biology, the RNA interference pathway is frequently utilized to study the role of genes in cell culture and in vivo in model organisms. With a sequence complementary to a gene of interest, double-stranded RNA is synthesized and inserted into a cell or organism, where it is recognized as exogenous genetic material and stimulates the pathway of RNAi. Researchers can cause a dramatic decrease in the expression of a targeted RNA gene by using this mechanism. The physiological function of the gene product can be seen by observing the effects of this decrease.
- Functional Genomics: Functional genomics using RNAi is particularly for genomic mapping.
- Medical: Potential antiviral therapies include topical microbicide treatments that use RNAi to treat the infection. RNA interference is also a promising way to treat cancers by silencing genes differentially upregulated in tumor cells or genes involved in cell division.
- Biotechnological Applications: RNA interference has been used for applications in biotechnology like food, insecticides, transgenic plants, etc.
References
- https://books.google.com/books?id=g3-ZSMossRYC
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3730950/
- https://eigenre.com/homepage/expert-perspectives/rnai/
- https://pubmed.ncbi.nlm.nih.gov/19021530
More To Read…
